PsPM
A matlab suite for Psycho Physiological Modelling
Current version: PsPM 6.1.0, released on 24.08.2023.
Introduction to PsPM
PsPM stands for PsychoPhysiological Modelling. It is a powerful matlab toolbox for model-based analysis of psychophysiological signals, for example SCR, ECG, respiration, pupil size, or startle eye-blink EMG. Currently, PsPM implements models for all of these modalities, and we are working towards further models, for example, for skin potential and ocular scan path length.
A PsPM allows inferring a psychological variable from observable physiological data. For example, associative memory can be inferred from observed skin conductance responses (SCR). This allows for quantitative description of hidden processes, increases the temporal resolution of analysis, and suppresses noise.
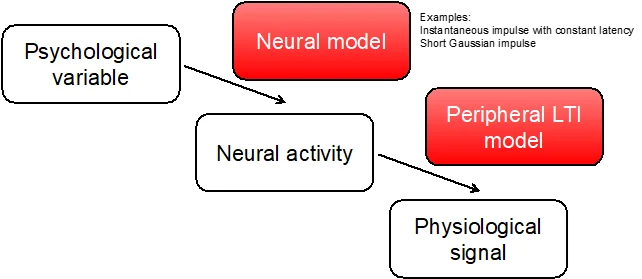
PsPM implements simple General Linear Convolution Models (GLM) for evoked SCR, or uses the Dynamic Causal Modelling (DCM) framework - as a tool to invert more complicated, non-linear models of SCR signals, for example for spontaneous fluctuations or anticipatory responses. Inference is drawn in a hierarchical summary-statistic approach (similar to SPM software for functional magnetic resonance imaging).
PsPM also supports other kinds of data for which no models exist yet, in particular we have extended support for eyetracking data.
The flexible software allows import of a number of data formats, including Spike, Biopac, VarioPort, (exported) ADInstruments LabChart, (exported) Biograph Infiniti, (exported) MindMedia BioTrace, Dataq/Windaq, AckKnowledge, ScanPhysLog, EDF, (exported) Eyelink, Matlab, and Text files.
Further features are simple programming of add-ons for import and modelling of new data types and automatic creation of batch scripts via the GUI.
PsPM incorporates the previous software package SCRalyze and offers all features of SCRalyze plus many more. If you started working on a project with SCRalyze and want to continue, you can still find previous software versions, help, and resources on http://scralyze.sourceforge.net.
PsPM is provided under the GNU General Public License (c) Dominik R. Bach, University of Bonn and University College London.
Learn more about the suite on our PsPM website.
PsPM courses
Course material
Course videocasts and slides are available on our PsPM website and on our channel on eLecture.
We offer regular PsPM courses. To check for updates, have a look on our news section, or follow bachlab on twitter/X.
Currently non scheduled.
Past PsPM courses
- 18 - 19 January 2024: University of Bonn (online 2-half-day PsPM satellite workshop with practical training)
- 21 - 22 July 2021 ESCAN 2021: Budapest (online 1.5-day PsPM satellite workshop with practical training)
- 30 June - 1 July 2020: ESCAN 2020 Budapest (full 2-day PsPM satellite workshop with practical training; postponed due to COVID-19 pandemic)
- 6 April - 14 May 2020: Live online course (see course recordings and slides below)
- 30 March 2020: EMHFC 2020 Bochum (workshop on pupil size measurements; cancelled due to COVID-19 pandemic)
- 13 November 2019: University of Göttingen
- 2 July 2019: Aegina Summer School on Social Cognition 2019
- 5 - 6 June 2019: Max Planck Institute for Human Cognitive and Brain Sciences Leipzig
- 29 - 30 May 2018: Pre-conference workshop, Psychology & Brain, Gießen
Course preparation
Getting started
- Make sure you have Matlab 2014 or higher
- Download the package
- Unzip it into any folder
- Put the folder on the matlab path using the path tool, or by typing addpath(‘folder’) s
- Start the GUI by typing “pspm” in the matlab command line
For the tutorials, we suggest that participants install Matlab and PsPM on their computers, download the provided data sets, and try to replicate the instructors’ demonstrations.
Further download material and useful links for all courses:
Latest PsPM version
Version PsPM 6.1.0, released on 24.08.2023.
Tutorial datasets
The tutorial text is part of the software manual. See subfolder “Manual” of the software download.
Help
- Help forum
- Bug reports
- Feature requests
- Sourceforge help forum (old versions until 4.2.0)
- Sourceforge bug reports (old versions until 4.1.0)


